ShapeWorks Studio Analyze Module
After optimization is complete, you may analyze the data in Studio using the Analysis module. There are several panels available in the Analysis Module.
View Panel
The View panel contains all of the options for controlling the analyis viewer options.
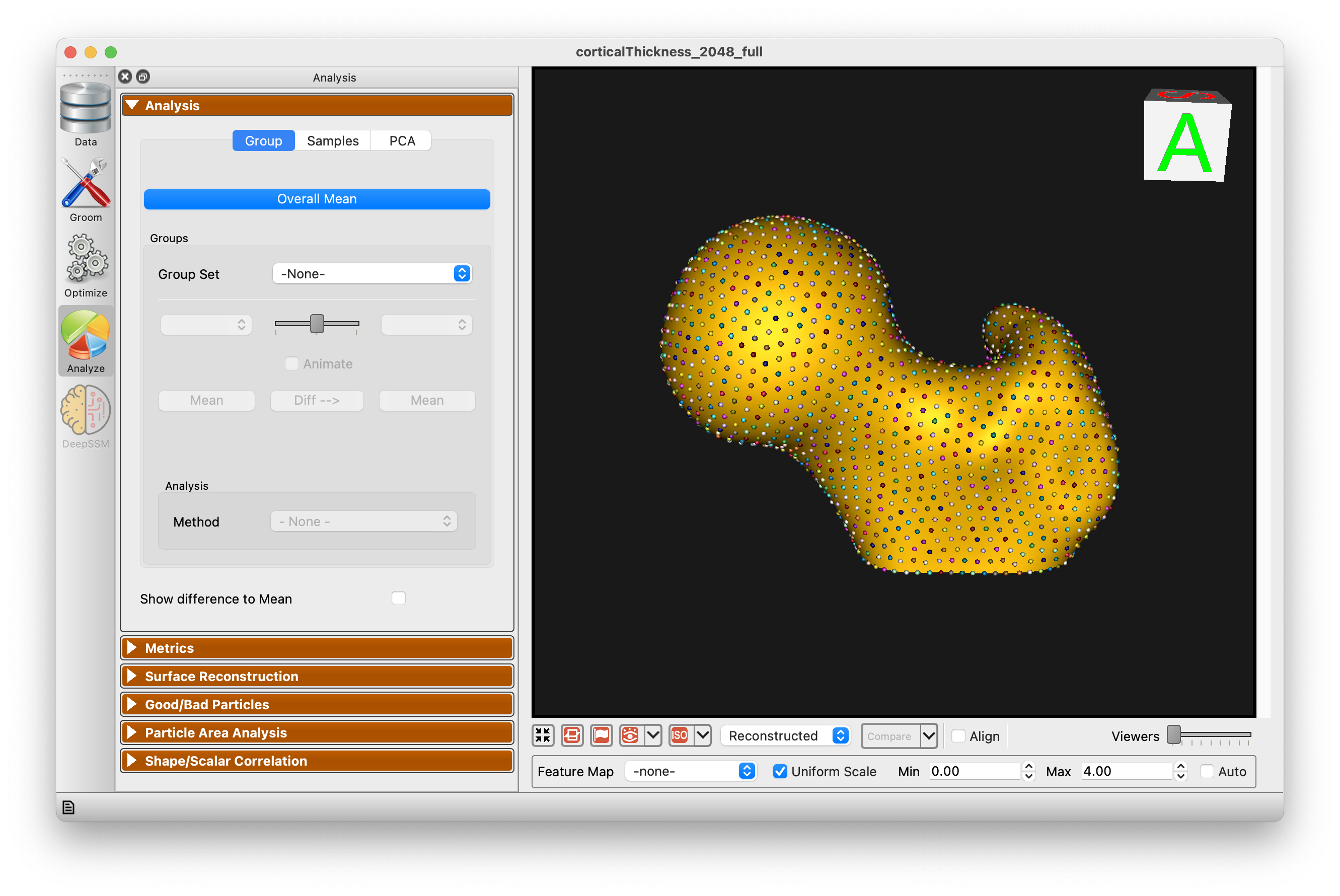
Group
The Group mode allows for group comparison to quantify and visualize population-level differences. Each group_<value> column from the project can be selected and the differences between any two values (i.e., group identifiers) in the groups can be displayed by selecting the group values in the left and right dropdown boxes. A group difference display showing vectors and a colormap of distance can be shown using the "Diff -->" button.
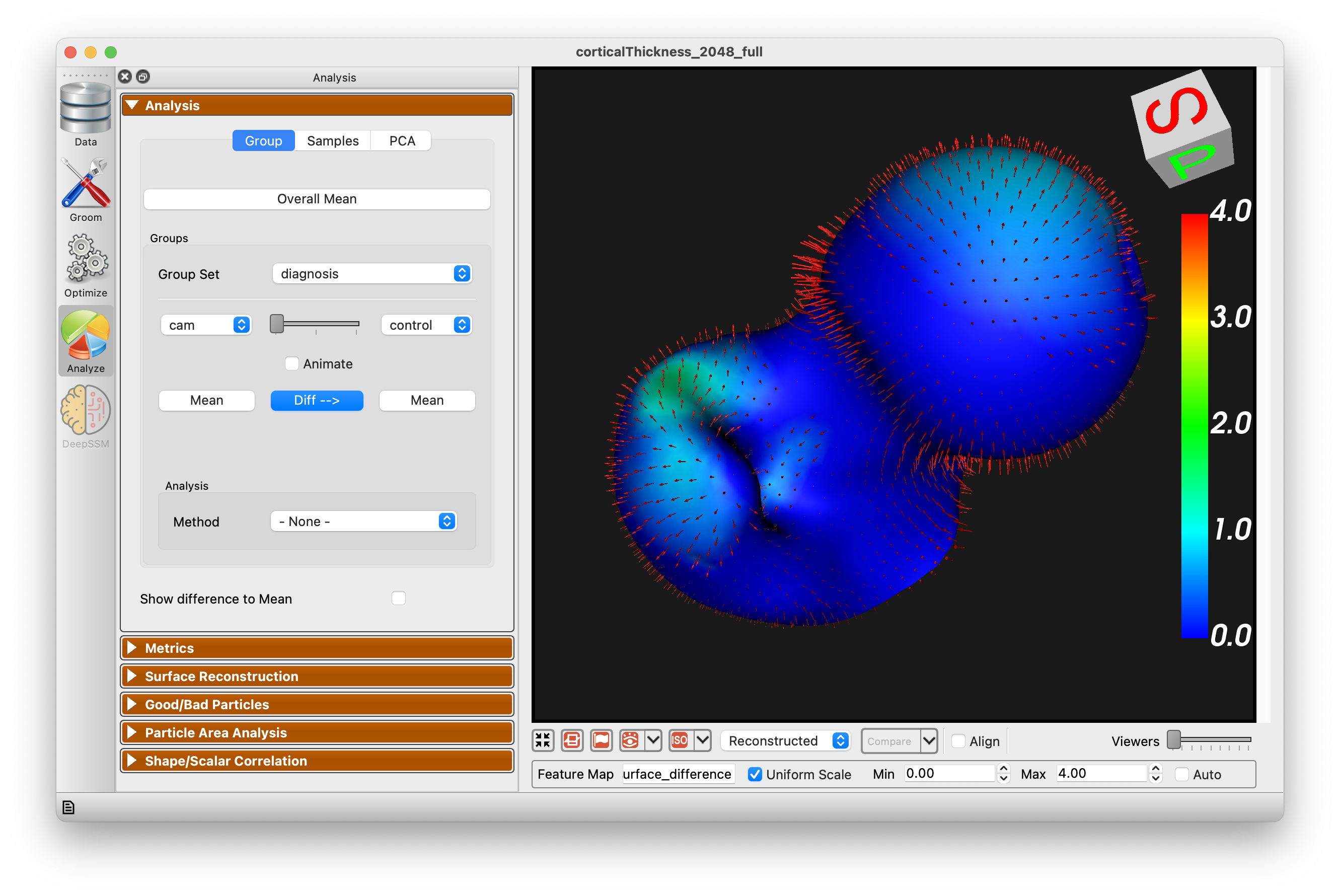
There are four different group analysis options available:
- LDA — Linear Discriminant Analysis finds the direction that best separates two groups by maximizing between-group variance relative to within-group variance.
- DWD — Distance Weighted Discrimination is a robust alternative to LDA that performs better in high-dimensional, small-sample settings common in shape modeling.
- Group Particle P-Values — Computes per-particle p-values between groups, highlighting regions of statistically significant shape difference.
- Network Analysis — Uses cluster-based statistical methods to identify spatially connected regions of significant group differences on the shape surface.
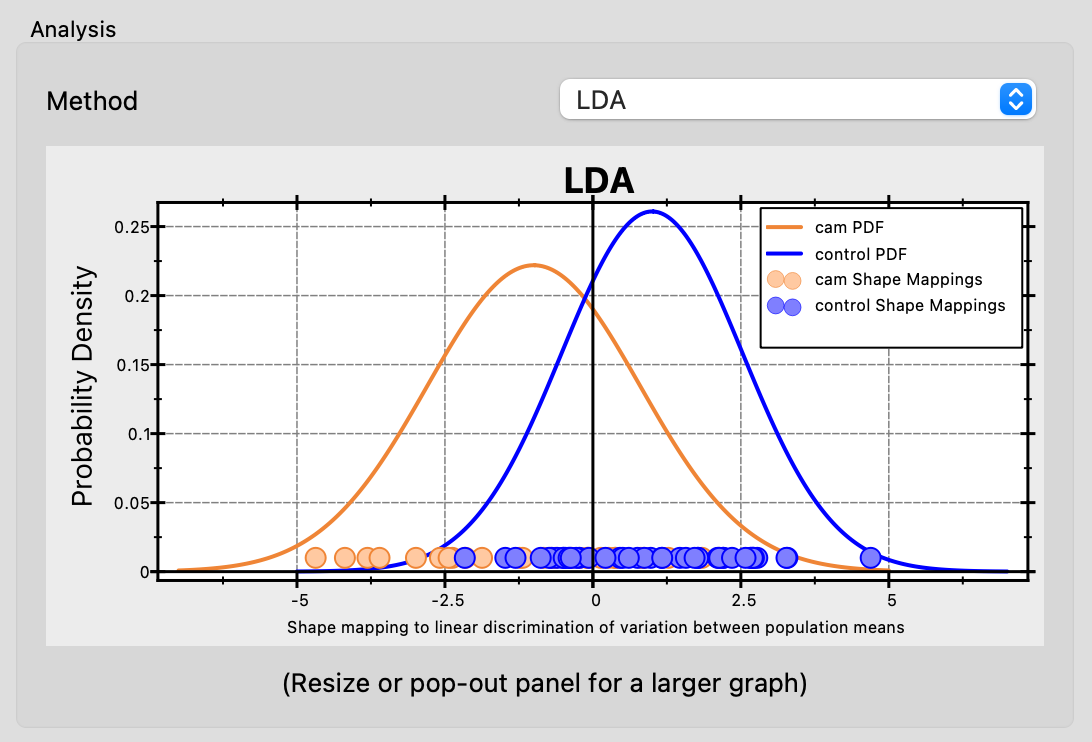
Both LDA and DWD produce a chart showing probability density functions (PDFs) for each group projected onto the discriminant direction, along with individual shape mappings as scatter points. This visualization helps assess how well the shape model distinguishes between populations.
LDA
Linear Discriminant Analysis (LDA) finds the linear direction in PCA space that best separates two groups by maximizing between-group variance relative to within-group variance.
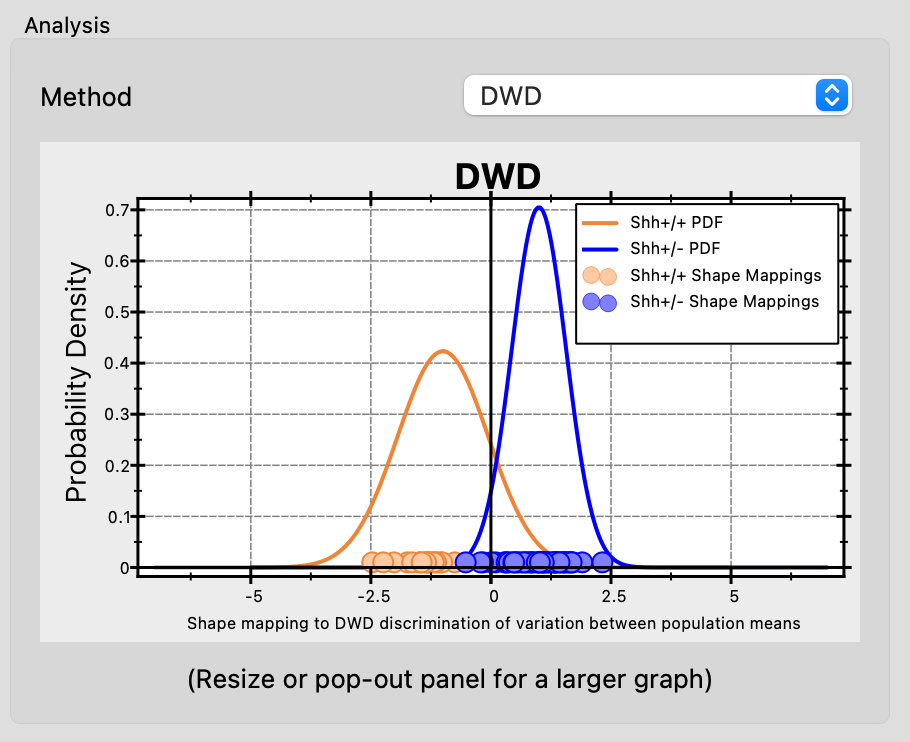
DWD
Distance Weighted Discrimination (DWD) finds a separating hyperplane that maximizes the average inverse distance of each sample to the boundary. This makes DWD more robust than LDA in high-dimensional, small-sample settings — a common scenario in shape modeling where the number of PCA modes can exceed the number of subjects. Each group requires at least 2 samples.
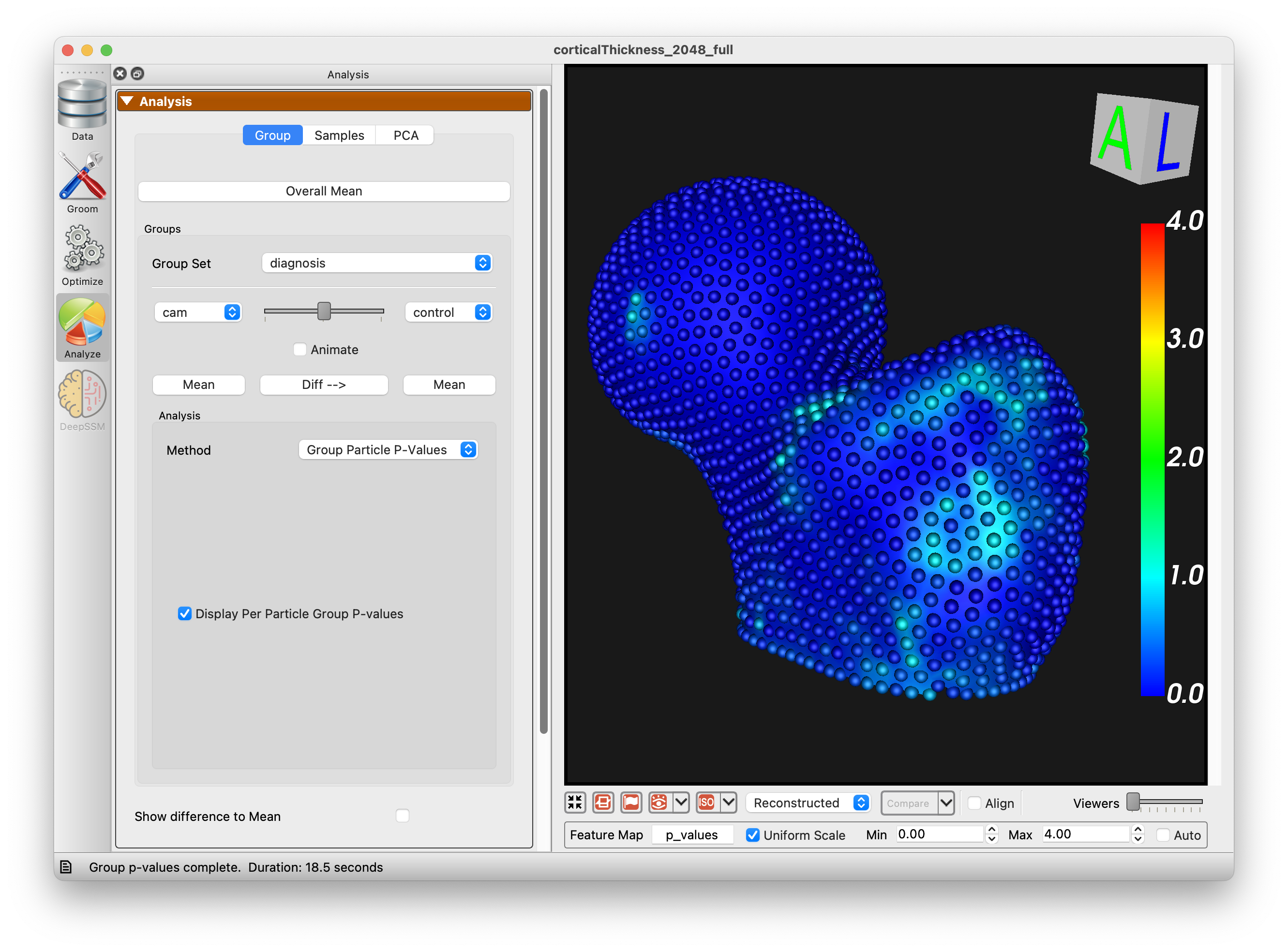
Group Particle P-Values
Computes a p-value for each correspondence particle to identify where two groups differ significantly in shape. For each particle, a Hotelling T-squared test compares the 3D positions across subjects in both groups. The test is repeated over multiple random subsamples (permutations) and the resulting p-values are corrected for multiple comparisons using false discovery rate (FDR) correction. Particles with low p-values indicate regions where the two groups have statistically significant shape differences. The p-values are displayed as a colormap on the shape surface.
Network Analysis
The Network Analysis tool provides a method to statistically analyze data captured in feature maps. Two implementations have been included: Network Analysis and SPM1D. Statistical parametric mapping (SPM, https://www.fil.ion.ucl.ac.uk/spm/) was introduced for the analysis of brain imaging data and has since been used in statistical analysis of surface-based group differences. The SPM1D option uses this technique without consideration of the connectivity and spatial relationship of the input data. The Network Analysis method uses the relative size of the network of connected correspondence particles to identify significant differences amongst groups, as originally described by Forman and colleagues (Forman SD, et al. Magnetic Resonance in Medicine33:636-647, 1995). Our publication on the specifics of this method and example applications is in review and will be referenced here when available.
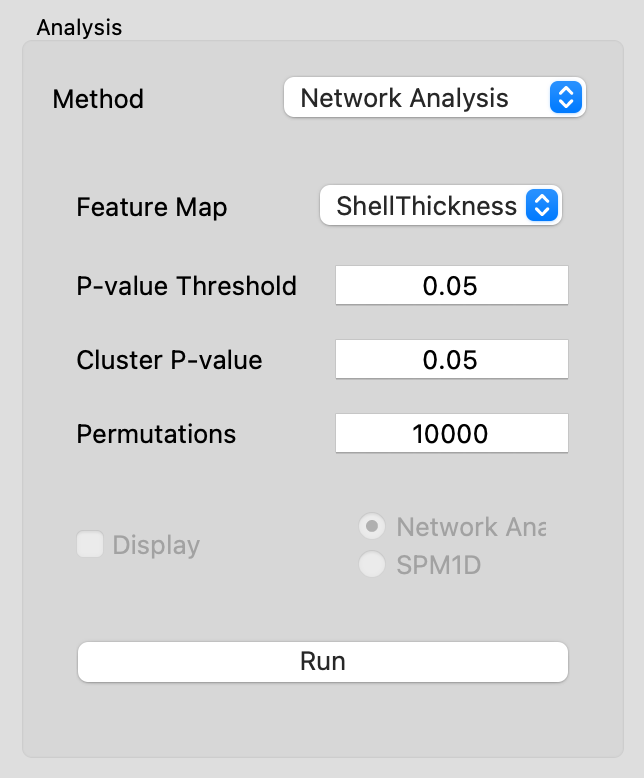
| Option | Description |
|---|---|
| Feature Map | Choose which feature to operate on. |
| P-value Threshold | The initial p-value used to threshold particles as an initial step to the evaluation of cluster size. Only modify from 0.05 for adjusted statistical assumptions. |
| Cluster P-value | The cluster p-value defines the p-value used to identify significant clusters for the group. Traditional statistics would assume a p-value of 0.05. |
| Permutations | This is the number of permutations used in the analysis. The maximum value is driven by the sample size, however 10,000 is generally considered acceptably large. |
| Display | Choose to display Network Analysis output or SPM1D |
After running the Network Analysis, you can enable the display of values with the "Display" checkbox:

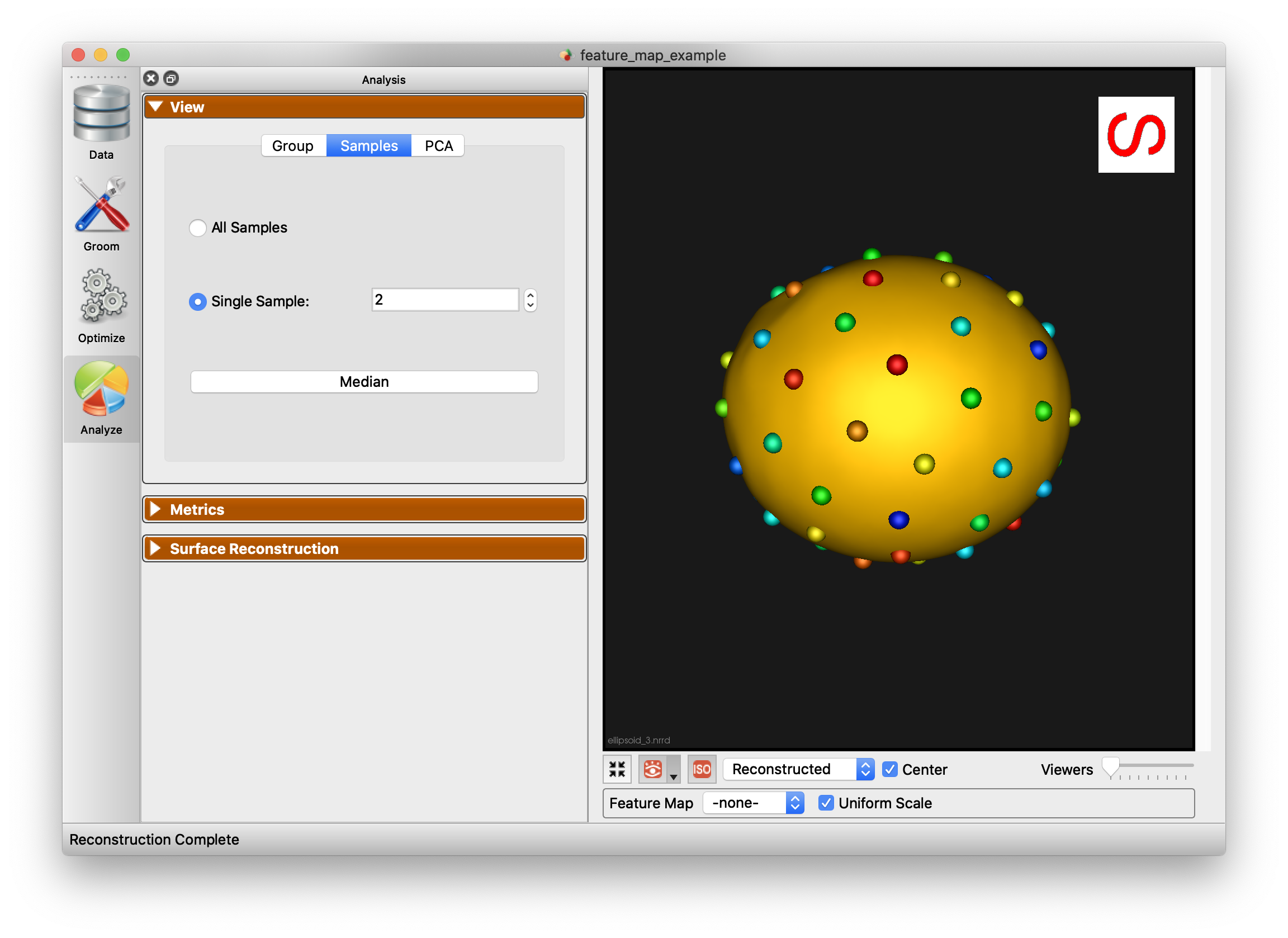
Samples
The Samples tab of the view panel allows you to view all of the samples in your cohort, view a single sample by index, or view the median sample.
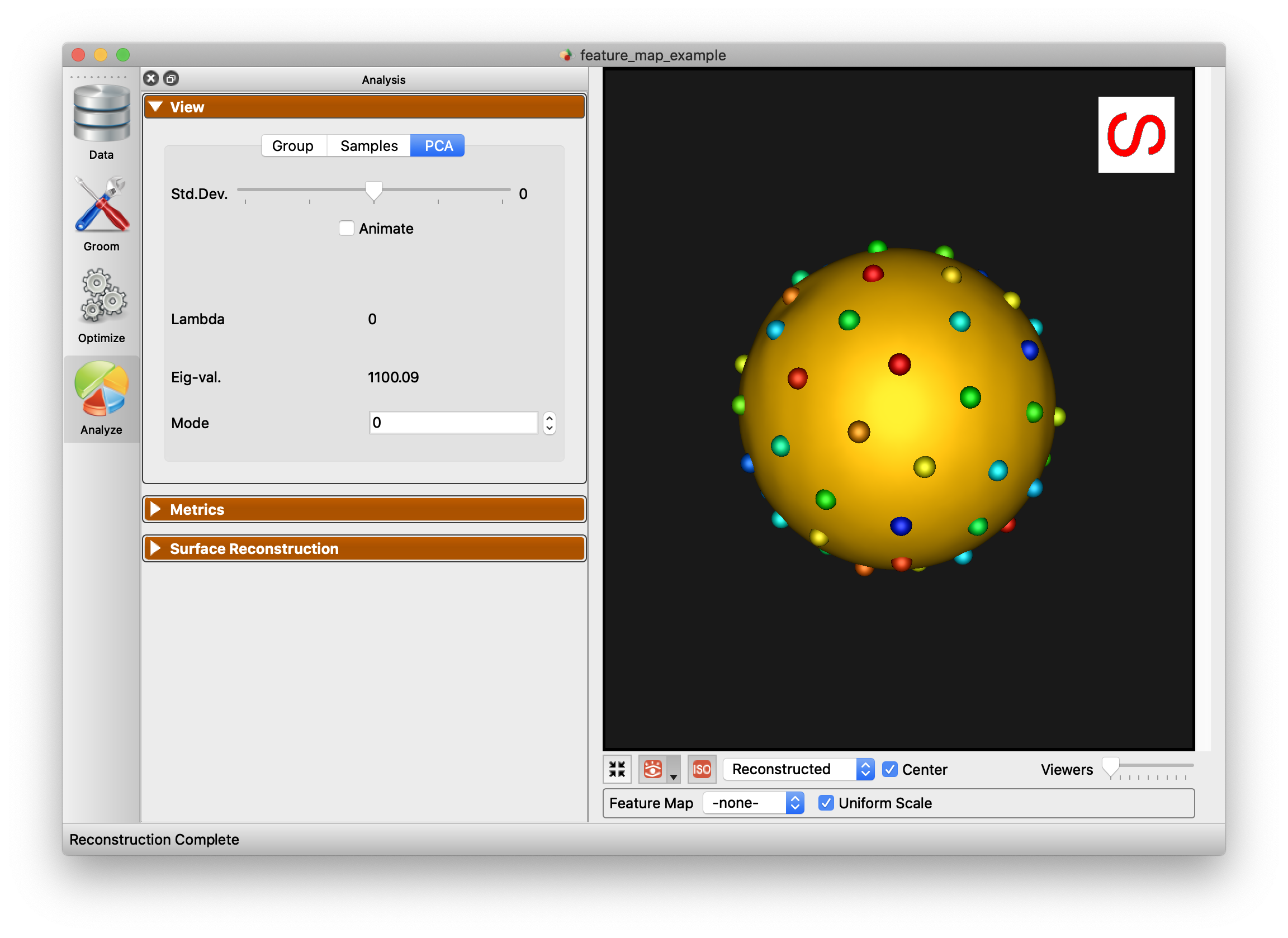
PCA
The PCA tab of the View panel shows reconstructed shapes (surface meshes) along PCA modes of variation. Modes of variations are shape parameters learned/discovered from your data that describes shape variations. You can control the location along the PCA mode with the Standard Deviation slider. The extent of this slider is configurable in Preferences. Values displayed:
Lambda - This shows the PCA loading of current position of the slider. The middle of the slider, at the mean value, will be 0. The extent of lambda is defined by the number of standard deviations of the slider as described above. At standard deviation of 1.0, it will be the square root of the mode's eigenvalue.
Eigenvalue - This shows the eigenvalue of the currently selected mode. This is a unitless measure of how much variance is explained by this mode. The modes are sorted in decreasing order of eigenvalue, corresponding to the most variance explained first.
Multi-Level PCA
The PCA tab of the View panel shows options to select modes of variation in different subspaces when a multiple domain shape model is loaded:
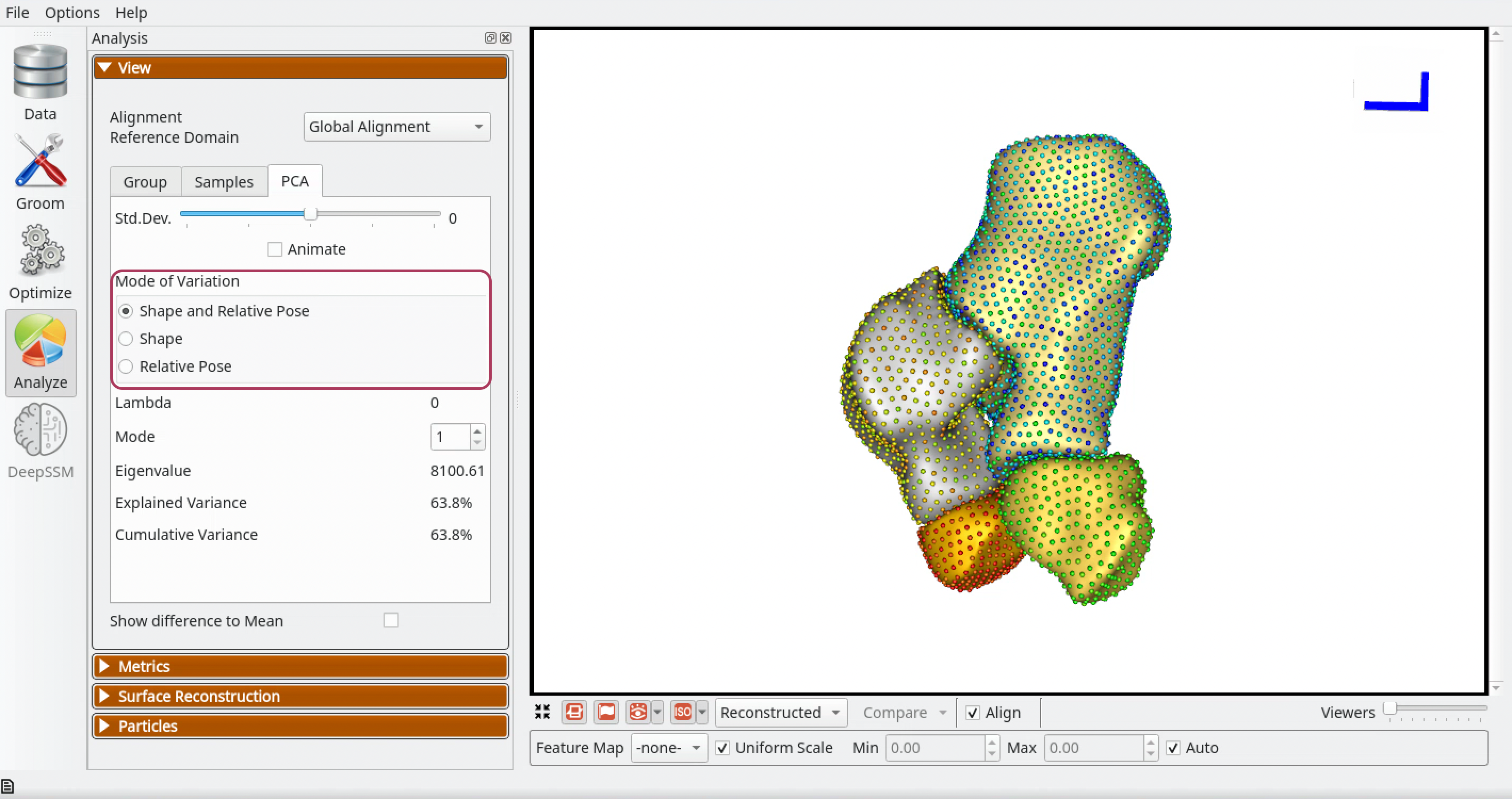
Shape and Relative Pose - Selecting this option shows reconstructed shapes and its eigenvalue and lambda along ordinary PCA modes of variation. PCA is done in the shared space of the multi-object shape structure and thus shape and pose variations are entangled here.
Shape - Selecting this option shows reconstructed shapes and its eigenvalue and lambda along morphological modes of variation. Multi-Level Component Analysis subtracts each domain's centroid per subject, removing translational pose differences. Note that rotational pose differences remain in the shape component.
Relative Pose - Selecting this option shows reconstructed shapes and its eigenvalue and lambda along relative pose modes of variation. Multi-Level Component Analysis keeps only per-domain centroids, showing translational relationships between domains. Note that rotational pose is not captured by this mode.
For a detailed explanation of these modes, their limitations, and how they interact with alignment settings, see Multi-Domain Reference Frames.
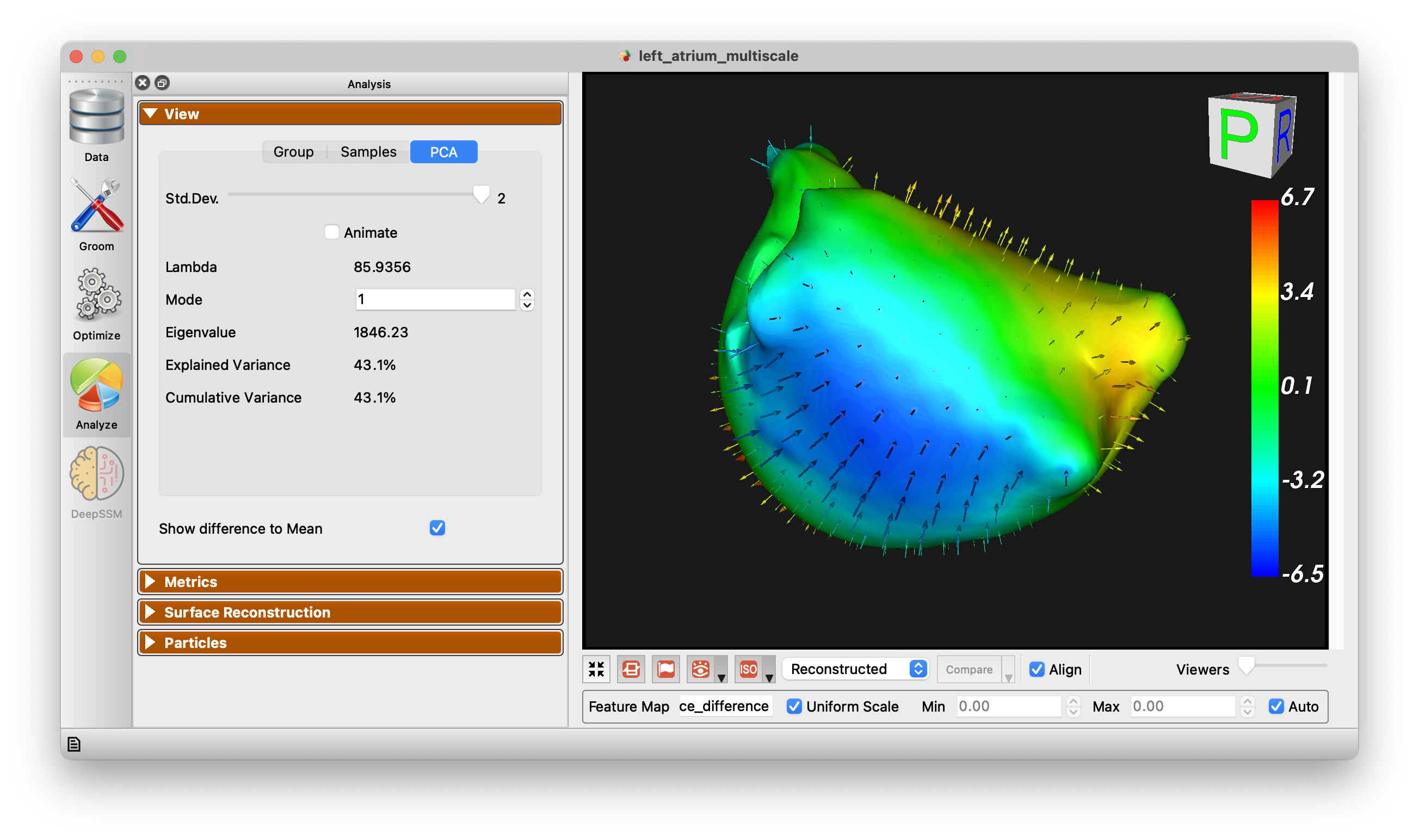
Show Difference to Mean
This checkbox allows for visualization from any surface to the mean. Arrow glyphs can be shown to display the difference as well.
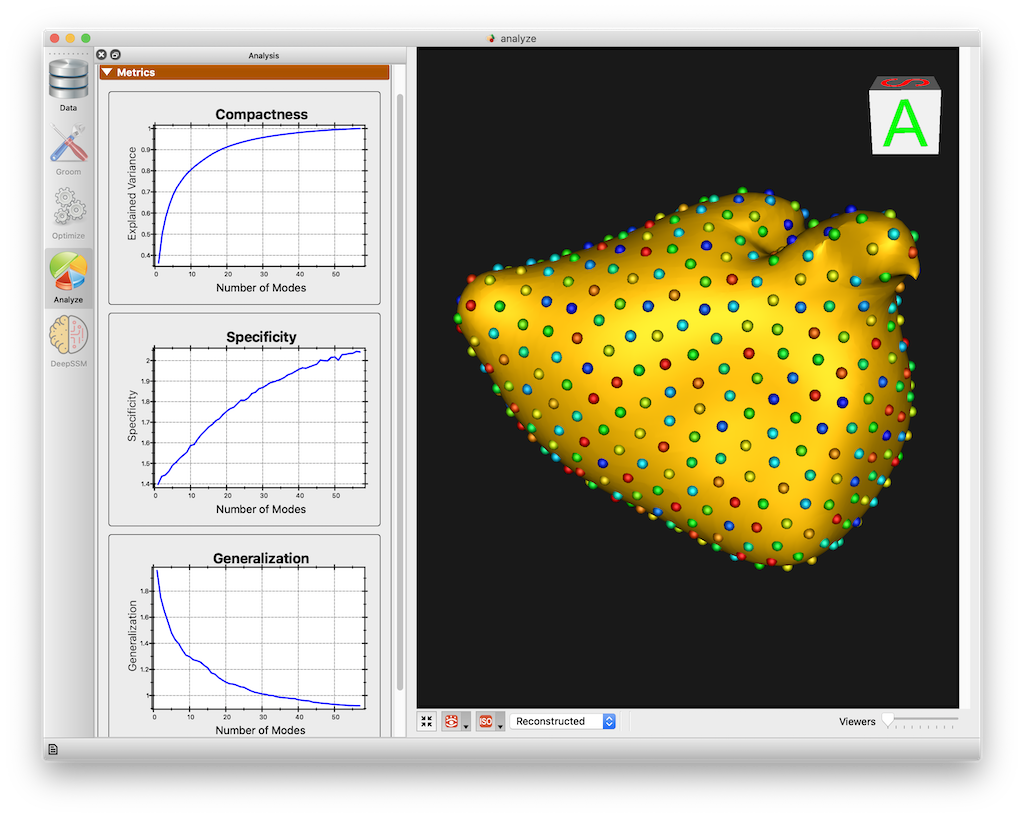
Metrics Panel
The Metrics panel shows the Compactness, Specificity, and Generalization charts. These are interactive charts with hover menus and the ability to zoom, pan, etc.
Distance Calculation Options
-
Particle to Particle (Default): Calculates the distance directly between particles when evaluating the metrics.
-
Particle to Surface: Calculates the distance between particles and the actual shape surface. This method enhances accuracy by measuring generalization as the distance between reconstructed particles and the true shape surface. Specificity is determined by the distance between sampled particles and the surface, ensuring generated shapes accurately reflect the true shape population and providing reliable and consistent metrics.
See Shape Model Evaluation for more information about shape evaluation.
Surface Reconstruction Panel
See the Surface Reconstruction page for details on reconstruction methods, warp method options, and examples.
Good/Bad Particles Panel
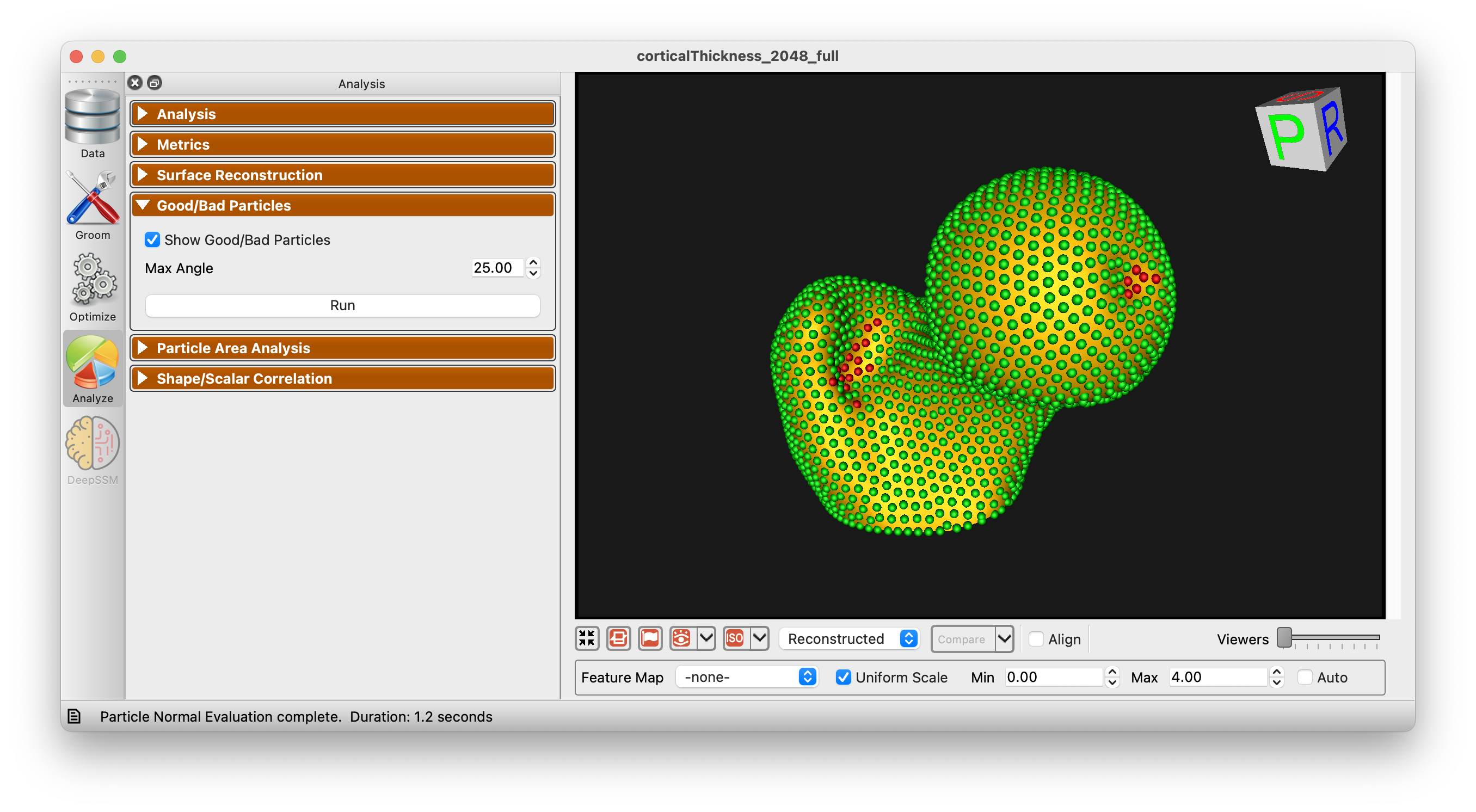
The Good/Bad Particles Panel enabled the display of "good/bad particles" in ShapeWorks Studio. A maximum angle parameter is given (default 45 degrees). When a particle's surface normal across subjects (averaged) exceeds this angle, the particle will be marked "bad" and shown in red. Good particles are shown in green. This can help in identifying problems with correspondence as it is typically expected that surface normals for given particles should be similar across subjects.
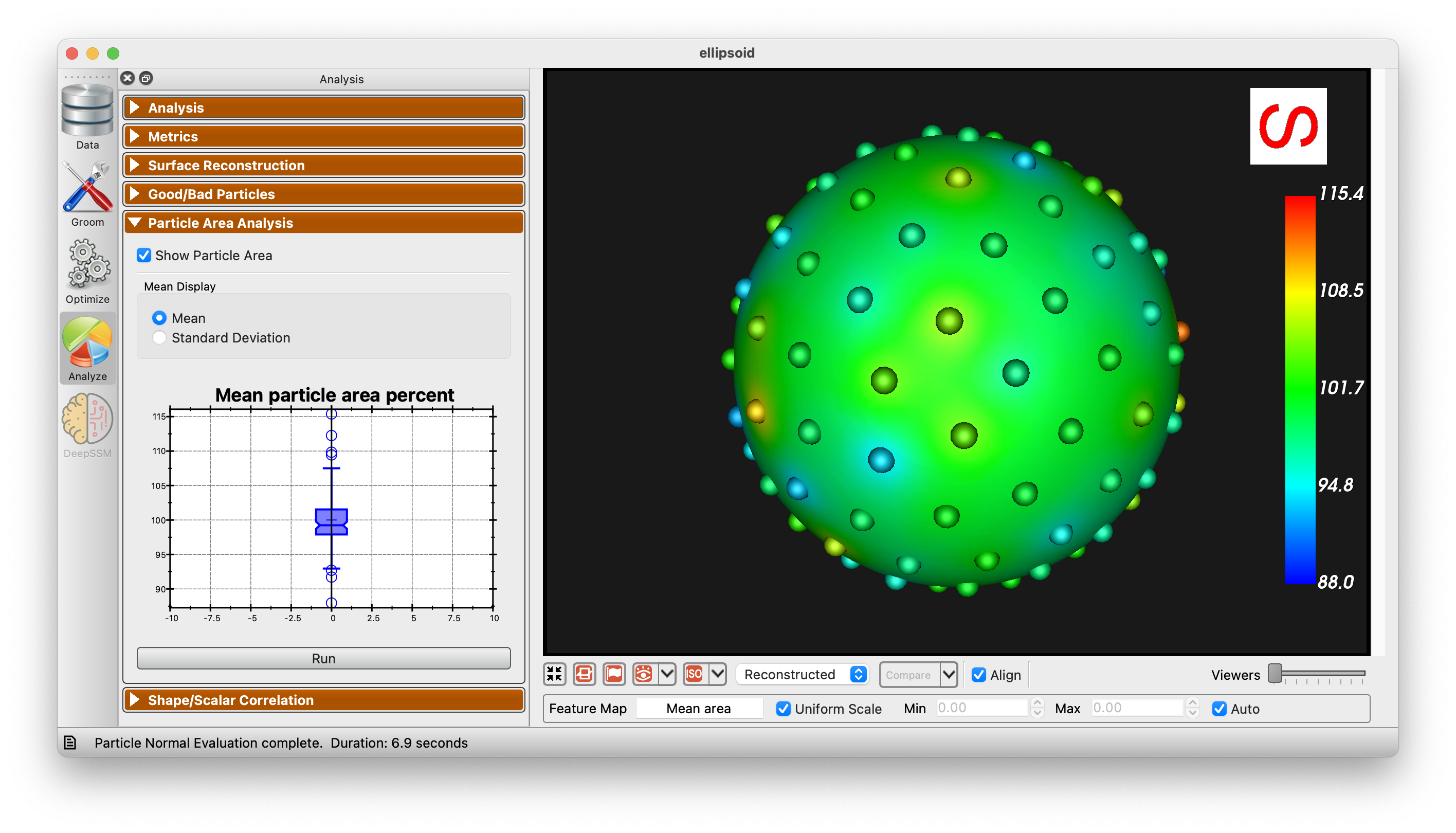
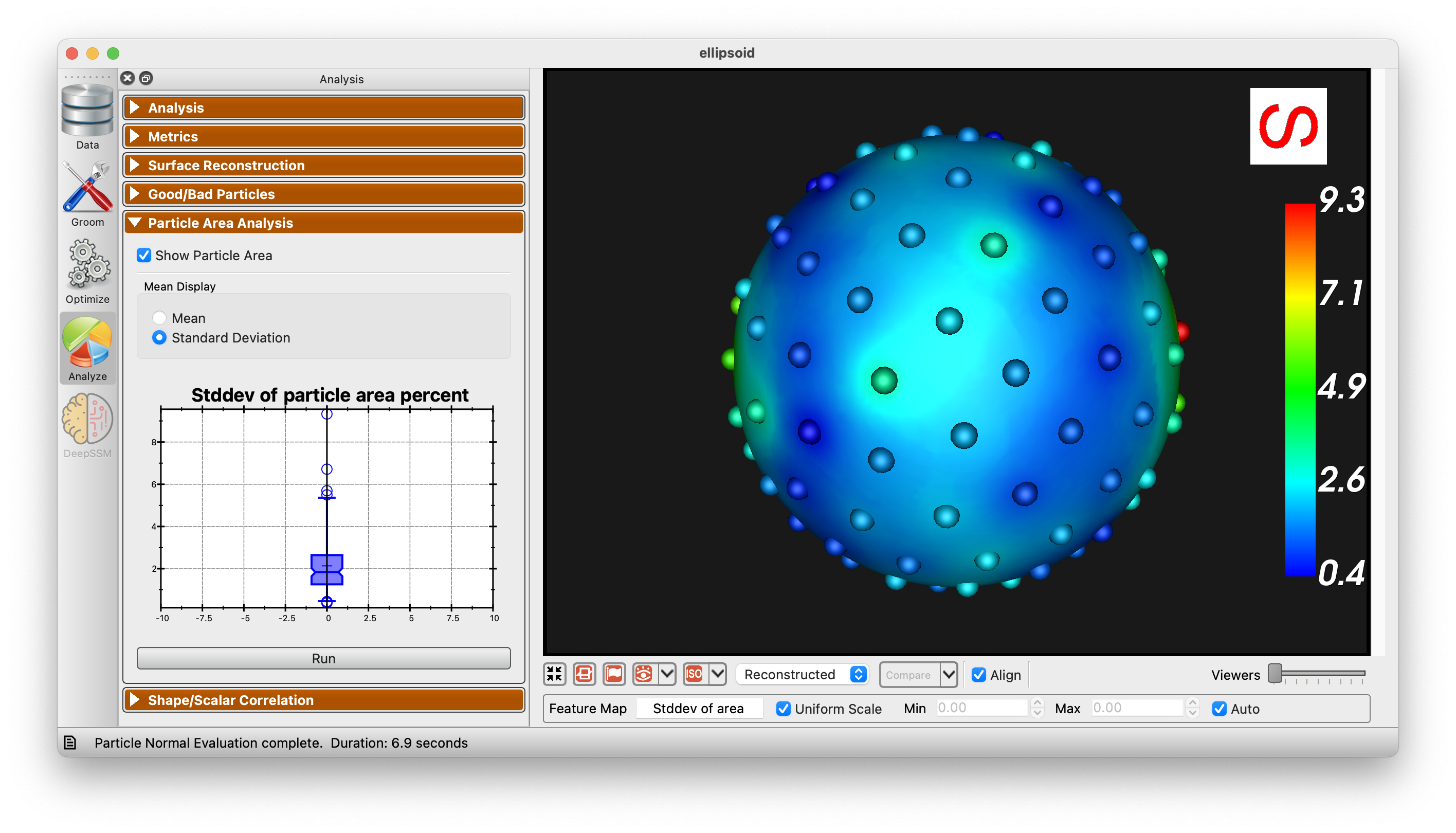
Particle Area Analysis
The Particle Area Analysis panel allows for the visualization of the area of each particle in the shape model. This can be useful for identifying areas that are not being captured well by the modell. There are two options for visualization, mean and standard deviation.
Shape/Scalar Correlation
The Shape/Scalar Correlation panel uses 2 block PLS regression to identify the relationship between shape and scalar data.
To use this panel, you must have scalar values (feature map). Choose the desired feature map from the dropdown menu.
Use the "Determine number of components" option to show the mean squared error for each number of components:
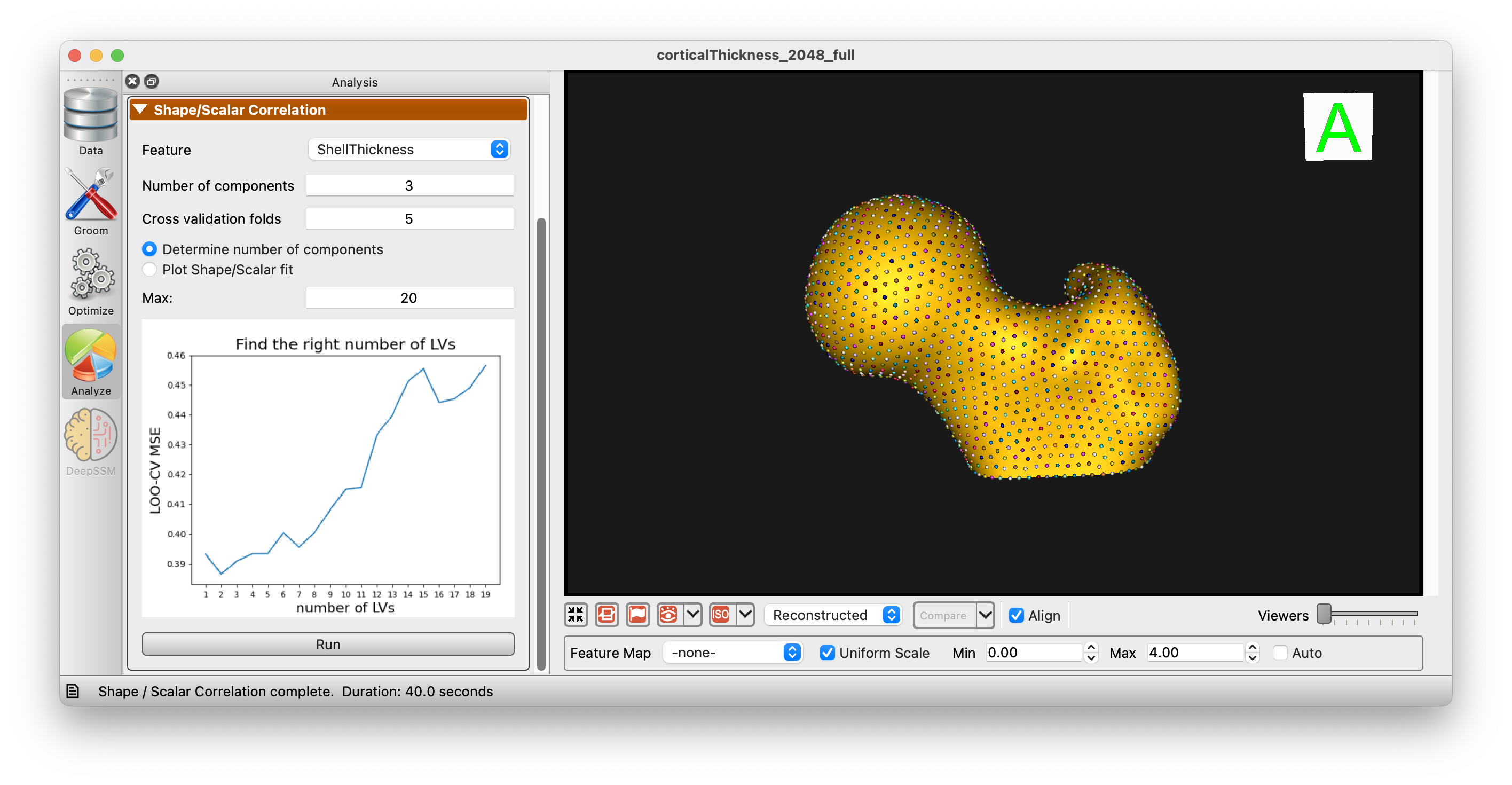
After choosing the desired number of components, you can plot the relationship between shape and scalar data:
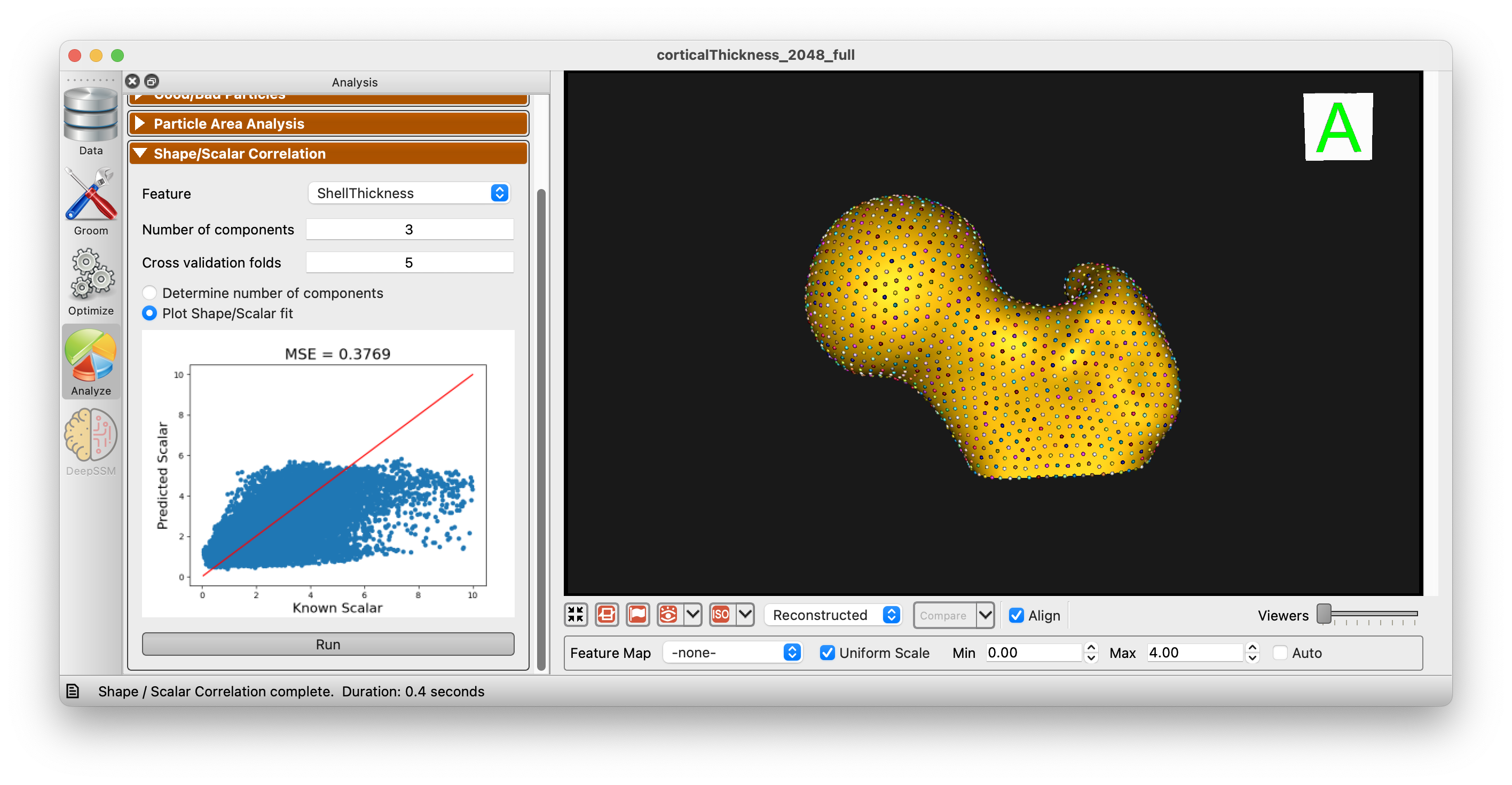
Additionally, you can use the 2 Block PLS regression to predict the scalar value from the shape data and vice versa:
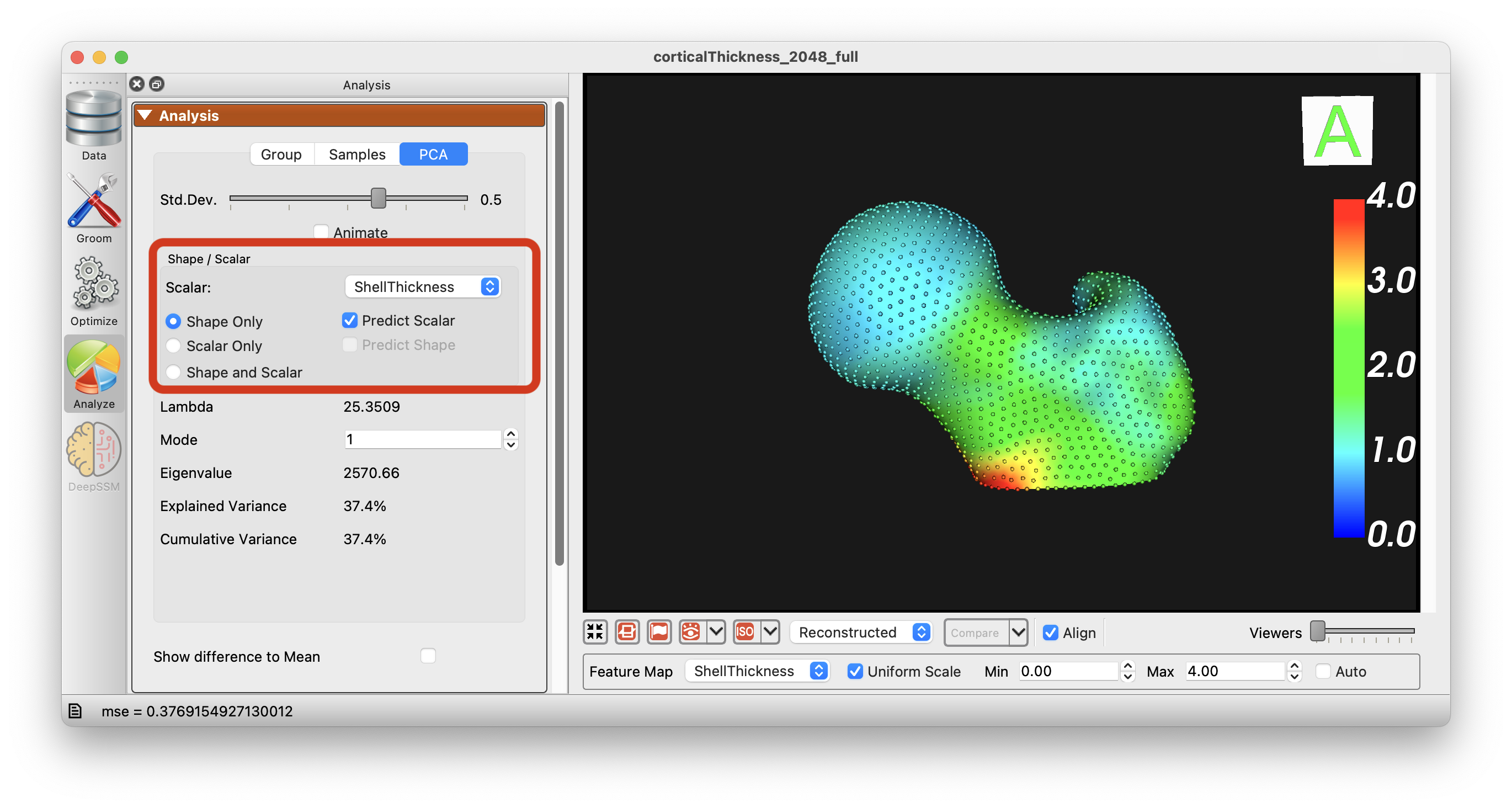
The options "Shape Only", "Scalar Only", and "Shape and Scalar" are available to choose from and the PCA will be performed accordingly.